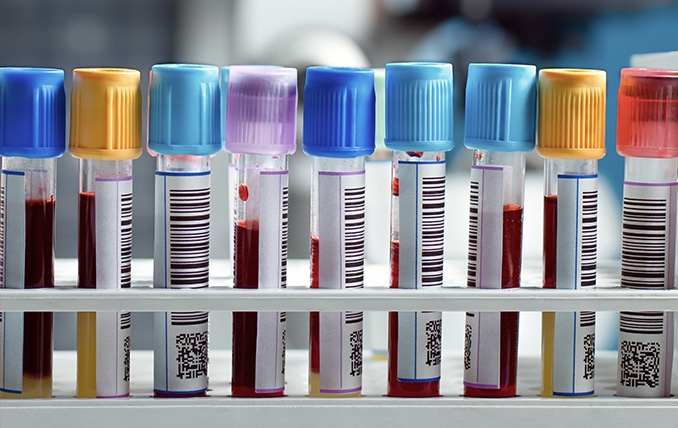
 Studying the real-time effects of labeling errors in the lab is extremely difficult. Billions of patient specimens, including blood and urine samples, as well as biopsies taken from multiple tissues and organs, are continually processed on a daily basis in clinical labs. Fortunately, several large-scale studies throughout the last 20 years have attempted to shed light on the clinical consequences of labeling errors in the clinic in an effort to improve patient care and reduce healthcare costs worldwide.
Studying the real-time effects of labeling errors in the lab is extremely difficult. Billions of patient specimens, including blood and urine samples, as well as biopsies taken from multiple tissues and organs, are continually processed on a daily basis in clinical labs. Fortunately, several large-scale studies throughout the last 20 years have attempted to shed light on the clinical consequences of labeling errors in the clinic in an effort to improve patient care and reduce healthcare costs worldwide.
The scale of mislabeling errors
It is nearly impossible to accurately measure the precise amount of medical errors that occur on a daily basis or the cost of labeling errors to patients and hospitals, but researchers have attempted to estimate the rate of errors at many institutions to gain a better understanding of the scope of mislabeling throughout the healthcare industry. In 1999, the US Institute of Medicine published a report that estimated the frequency of medical errors resulting in patient death to between 44,000 and 98,000 per year in the United States. Since then, studies began investigating large numbers of medical institutions, finding mislabeling rates ranging from 0.139 to 7.4 errors per 1000 specimens, depending on the hospital and the unit. In Canada, an inquiry into the mislabeling of samples from patients with breast cancer found that 42% out of the tumor samples that initially tested negative for estrogen receptor (ER)/progesterone receptor (PR) were ER/PR positive upon retesting. Had these patients not been retested, they would not have received hormonal therapy, which has proven to be of particular benefit for ER/PR-positive patients.
The consequences of clinical labeling errors
Specimen labeling errors have repercussions for both the patient and the institution providing care. The two most important consequences of mislabeled specimens are the failure to provide proper and immediate care and the provision of inappropriate care.1 Not receiving the necessary treatment or receiving the wrong treatment can harm the patient, resulting in unnecessary morbidity and mortality. One study found that mislabeling errors in blood transfusions caused 40% to 50% of all transfusion-related morbidities.4 Patients may also suffer additional stress from receiving the wrong diagnosis or from having to wait a longer period of time to receive a diagnosis after undergoing repeat testing. Repeat testing, particularly tumor biopsies, expose the patient to unnecessary risks and complications; even blood draws can be uncomfortable and, when performed too frequently, can lead to anemia and worsen the patient’s condition.
The financial cost of mislabeling for patients and the healthcare system as a whole can be extensive. The College of Pathologists evaluated the average cost of labeling errors at approximately $712 per specimen. By multiplying this cost by the number of mislabeled specimens, they estimate that it costs hospitals and other healthcare institutions $280,000 per million specimens, adding up to over $1 million dollars a year for some of the larger institutes. These costs are directly associated with redrawing of the specimen—including labor, consumables, and supply costs—reanalysis, additional nurse and physician costs, and prolonged hospital stays.3 The number and severity of the errors can also reflect poorly on the healthcare institution itself, damaging its reputation, and, over time, can erode overall confidence in the healthcare system.4
Labeling errors in a non-clinical lab can prove to be costly in terms of time, money, and effort. However, labeling errors in the clinic can have such dire consequences. When dealing with patient samples, the utmost care must be taken to ensure no errors are introduced in the analysis process, which can lead to misdiagnosis and the implementation of inappropriate treatment options. In many different ways, labeling can go wrong, using the right label for the application becomes crucial. Come back next week for the following blog in this series, for some of the commonly used solutions to address labeling challenges in the clinic.
LabTAG by GA International is a leading manufacturer of high-performance specialty labels and a supplier of identification solutions used in research and medical labs as well as healthcare institutions.
References:
- Kahn SE. Specimen Mislabeling: A Significant and Costly Cause of Potentially Serious Medical Errors. Brønshøj, Denmark; 2005.
- Kim JK, Dotson B, Thomas S, Nelson KC. Standardized patient identification and specimen labeling: A retrospective analysis on improving patient safety. J Am Acad Dermatol. 2013. doi:10.1016/j.jaad.2012.06.017
- The Problem of Mislabeled Specimens. Northfield, IL; 2010.
- Martin, H; Metcalfe, S; Whichello R. Specimen Labeling Errors: A Retrospective Study. Online J Nurs Informatics. 2015;19(2):1-11.


